Journal Description
Journal of Developmental Biology
Journal of Developmental Biology
is a peer-reviewed, open access journal on the development of multicellular organisms at the molecule, cell, tissue, organ and whole organism levels, and is published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), PubMed, PMC, PubAg, CAPlus / SciFinder, and other databases.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 17.1 days after submission; acceptance to publication is undertaken in 3.7 days (median values for papers published in this journal in the first half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about Journal of Developmental Biology.
Impact Factor:
2.7 (2022);
5-Year Impact Factor:
3.0 (2022)
Latest Articles
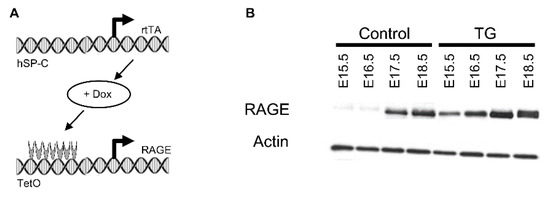
Decreased Expression of Pulmonary Homeobox NKX2.1 and Surfactant Protein C in Developing Lungs That Over-Express Receptors for Advanced Glycation End-Products (RAGE)
J. Dev. Biol. 2023, 11(3), 33; https://doi.org/10.3390/jdb11030033 - 15 Jul 2023
Abstract
►
Show Figures
Receptors for advanced glycation end-products (RAGE) are multi-ligand cell surface receptors of the immunoglobin superfamily prominently expressed by lung epithelium. Previous experiments demonstrated that over-expression of RAGE by murine alveolar epithelium throughout embryonic development causes neonatal lethality coincident with significant lung hypoplasia. In
[...] Read more.
Receptors for advanced glycation end-products (RAGE) are multi-ligand cell surface receptors of the immunoglobin superfamily prominently expressed by lung epithelium. Previous experiments demonstrated that over-expression of RAGE by murine alveolar epithelium throughout embryonic development causes neonatal lethality coincident with significant lung hypoplasia. In the current study, we evaluated the expression of NKX2.1 (also referred to as TTF-1), a homeodomain-containing transcription factor critical for branching morphogenesis, in mice that differentially expressed RAGE. We also contextualized NKX2.1 expression with the abundance of FoxA2, a winged double helix DNA binding protein that influences respiratory epithelial cell differentiation and surfactant protein expression. Conditional RAGE over-expression was induced in mouse lung throughout gestation (embryonic day E0–18.5), as well as during the critical saccular period of development (E15.5–18.5), and analyses were conducted at E18.5. Histology revealed markedly less lung parenchyma beginning in the canalicular stage of lung development and continuing throughout the saccular period. We discovered consistently decreased expression of both NKX2.1 and FoxA2 in lungs from transgenic (TG) mice compared to littermate controls. We also observed diminished surfactant protein C in TG mice, suggesting possible hindered differentiation and/or proliferation of alveolar epithelial cells under the genetic control of these two critical transcription factors. These results demonstrate that RAGE must be specifically regulated during lung formation. Perturbation of epithelial cell differentiation culminating in respiratory distress and perinatal lethality may coincide with elevated RAGE expression in the lung parenchyma.
Full article
Open AccessReview
Evolutionary Change in Gut Specification in Caenorhabditis Centers on the GATA Factor ELT-3 in an Example of Developmental System Drift
J. Dev. Biol. 2023, 11(3), 32; https://doi.org/10.3390/jdb11030032 - 08 Jul 2023
Abstract
Cells in a developing animal embryo become specified by the activation of cell-type-specific gene regulatory networks. The network that specifies the gut in the nematode Caenorhabditis elegans has been the subject of study for more than two decades. In this network, the maternal
[...] Read more.
Cells in a developing animal embryo become specified by the activation of cell-type-specific gene regulatory networks. The network that specifies the gut in the nematode Caenorhabditis elegans has been the subject of study for more than two decades. In this network, the maternal factors SKN-1/Nrf and POP-1/TCF activate a zygotic GATA factor cascade consisting of the regulators MED-1,2 → END-1,3 → ELT-2,7, leading to the specification of the gut in early embryos. Paradoxically, the MED, END, and ELT-7 regulators are present only in species closely related to C. elegans, raising the question of how the gut can be specified without them. Recent work found that ELT-3, a GATA factor without an endodermal role in C. elegans, acts in a simpler ELT-3 → ELT-2 network to specify gut in more distant species. The simpler ELT-3 → ELT-2 network may thus represent an ancestral pathway. In this review, we describe the elucidation of the gut specification network in C. elegans and related species and propose a model by which the more complex network might have formed. Because the evolution of this network occurred without a change in phenotype, it is an example of the phenomenon of Developmental System Drift.
Full article
(This article belongs to the Special Issue The 10th Anniversary of JDB: Feature Papers)
►▼
Show Figures
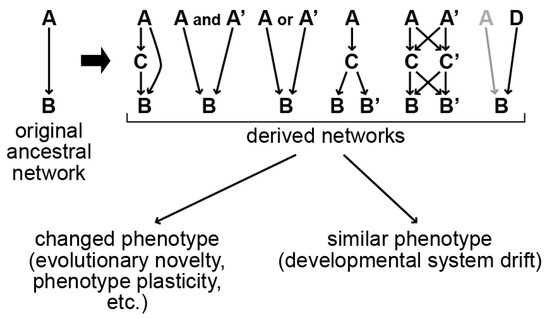
Figure 1
Open AccessBrief Report
Patterning of the Vertebrate Head in Time and Space by BMP Signaling
J. Dev. Biol. 2023, 11(3), 31; https://doi.org/10.3390/jdb11030031 - 03 Jul 2023
Abstract
How head patterning is regulated in vertebrates is yet to be understood. In this study, we show that frog embryos injected with Noggin at different blastula and gastrula stages had their head development sequentially arrested at different positions. When timed BMP inhibition was
[...] Read more.
How head patterning is regulated in vertebrates is yet to be understood. In this study, we show that frog embryos injected with Noggin at different blastula and gastrula stages had their head development sequentially arrested at different positions. When timed BMP inhibition was applied to BMP-overexpressing embryos, the expression of five genes: xcg-1 (a marker of the cement gland, which is the front-most structure in the frog embryo), six3 (a forebrain marker), otx2 (a forebrain and mid-brain marker), gbx2 (an anterior hindbrain marker), and hoxd1 (a posterior hindbrain marker) were sequentially fixed. These results suggest that the vertebrate head is patterned from anterior to posterior in a progressive fashion and may involve timed actions of the BMP signaling.
Full article
(This article belongs to the Special Issue 2022 Feature Papers by JDB’s Editorial Board Members)
►▼
Show Figures

Figure 1
Open AccessFeature PaperArticle
Molecular and Cellular Characterization of Avian Reticulate Scales Implies the Evo–Devo Novelty of Skin Appendages in Foot Sole
J. Dev. Biol. 2023, 11(3), 30; https://doi.org/10.3390/jdb11030030 - 03 Jul 2023
Abstract
Among amniotic skin appendages, avian feathers and mammalian hairs protect their stem cells in specialized niches, located in the collar bulge and hair bulge, respectively. In chickens and alligators, label retaining cells (LRCs), which are putative stem cells, are distributed in the hinge
[...] Read more.
Among amniotic skin appendages, avian feathers and mammalian hairs protect their stem cells in specialized niches, located in the collar bulge and hair bulge, respectively. In chickens and alligators, label retaining cells (LRCs), which are putative stem cells, are distributed in the hinge regions of both avian scutate scales and reptilian overlapping scales. These LRCs take part in scale regeneration. However, it is unknown whether other types of scales, for example, symmetrically shaped reticulate scales, have a similar way of preserving their stem cells. In particular, the foot sole represents a special interface between animal feet and external environments, with heavy mechanical loading. This is different from scutate-scale-covered metatarsal feet that function as protection. Avian reticulate scales on foot soles display specialized characteristics in development. They do not have a placode stage and lack β-keratin expression. Here, we explore the molecular and cellular characteristics of avian reticulate scales. RNAscope analysis reveals different molecular profiles during surface and hinge determination compared with scutate scales. Furthermore, reticulate scales express Keratin 15 (K15) sporadically in both surface- and hinge-region basal layer cells, and LRCs are not localized. Upon wounding, the reticulate scale region undergoes repair but does not regenerate. Our results suggest that successful skin appendage regeneration requires localized stem cell niches to guide regeneration.
Full article
(This article belongs to the Special Issue Development of the Skin in Vertebrates)
►▼
Show Figures
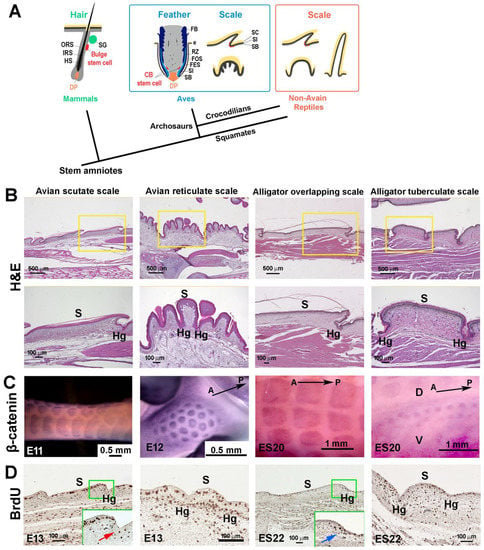
Figure 1
Open AccessArticle
The Tumor Suppressor Adenomatous Polyposis Coli (apc) Is Required for Neural Crest-Dependent Craniofacial Development in Zebrafish
by
, , , , , , , , , , and
J. Dev. Biol. 2023, 11(3), 29; https://doi.org/10.3390/jdb11030029 - 29 Jun 2023
Abstract
Neural crest (NC) is a unique vertebrate cell type arising from the border of the neural plate and epidermis that gives rise to diverse tissues along the entire body axis. Roberto Mayor and colleagues have made major contributions to our understanding of NC
[...] Read more.
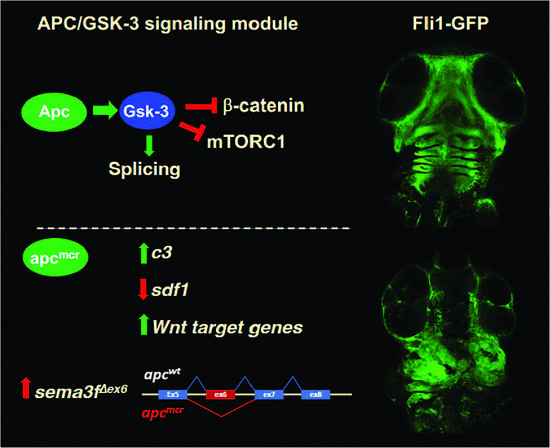
Neural crest (NC) is a unique vertebrate cell type arising from the border of the neural plate and epidermis that gives rise to diverse tissues along the entire body axis. Roberto Mayor and colleagues have made major contributions to our understanding of NC induction, delamination, and migration. We report that a truncating mutation of the classical tumor suppressor Adenomatous Polyposis Coli (apc) disrupts craniofacial development in zebrafish larvae, with a marked reduction in the cranial neural crest (CNC) cells that contribute to mandibular and hyoid pharyngeal arches. While the mechanism is not yet clear, the altered expression of signaling molecules that guide CNC migration could underlie this phenotype. For example, apcmcr/mcr larvae express substantially higher levels of complement c3, which Mayor and colleagues showed impairs CNC cell migration when overexpressed. However, we also observe reduction in stroma-derived factor 1 (sdf1/cxcl12), which is required for CNC migration into the head. Consistent with our previous work showing that APC directly enhances the activity of glycogen synthase kinase 3 (GSK-3) and, independently, that GSK-3 phosphorylates multiple core mRNA splicing factors, we identify 340 mRNA splicing variations in apc mutant zebrafish, including a splice variant that deletes a conserved domain in semaphorin 3f (sema3f), an axonal guidance molecule and a known regulator of CNC migration. Here, we discuss potential roles for apc in CNC development in the context of some of the seminal findings of Mayor and colleagues.
Full article
(This article belongs to the Special Issue From Cell to Embryo: A Theme Issue Honoring Professor Dr. Roberto Mayor)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Regionalized Protein Localization Domains in the Zebrafish Hair Cell Kinocilium
by
, , , and
J. Dev. Biol. 2023, 11(2), 28; https://doi.org/10.3390/jdb11020028 - 16 Jun 2023
Abstract
Sensory hair cells are the receptors for auditory, vestibular, and lateral line sensory organs in vertebrates. These cells are distinguished by “hair”-like projections from their apical surface collectively known as the hair bundle. Along with the staircase arrangement of the actin-filled stereocilia, the
[...] Read more.
Sensory hair cells are the receptors for auditory, vestibular, and lateral line sensory organs in vertebrates. These cells are distinguished by “hair”-like projections from their apical surface collectively known as the hair bundle. Along with the staircase arrangement of the actin-filled stereocilia, the hair bundle features a single, non-motile, true cilium called the kinocilium. The kinocilium plays an important role in bundle development and the mechanics of sensory detection. To understand more about kinocilial development and structure, we performed a transcriptomic analysis of zebrafish hair cells to identify cilia-associated genes that have yet to be characterized in hair cells. In this study, we focused on three such genes—ankef1a, odf3l2a, and saxo2—because human or mouse orthologs are either associated with sensorineural hearing loss or are located near uncharacterized deafness loci. We made transgenic fish that express fluorescently tagged versions of their proteins, demonstrating their localization to the kinocilia of zebrafish hair cells. Furthermore, we found that Ankef1a, Odf3l2a, and Saxo2 exhibit distinct localization patterns along the length of the kinocilium and within the cell body. Lastly, we have reported a novel overexpression phenotype of Saxo2. Overall, these results suggest that the hair cell kinocilium in zebrafish is regionalized along its proximal-distal axis and set the groundwork to understand more about the roles of these kinocilial proteins in hair cells.
Full article
(This article belongs to the Special Issue Zebrafish—a Model System for Developmental Biology II)
►▼
Show Figures
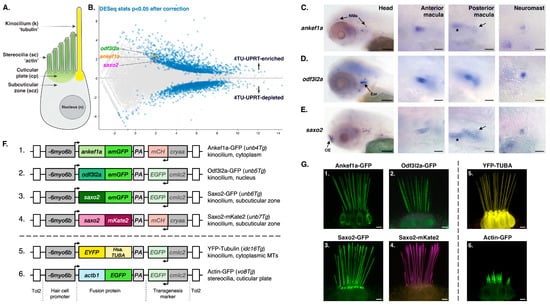
Figure 1
Open AccessFeature PaperReview
The Lost and Found: Unraveling the Functions of Orphan Genes
J. Dev. Biol. 2023, 11(2), 27; https://doi.org/10.3390/jdb11020027 - 13 Jun 2023
Abstract
Orphan Genes (OGs) are a mysterious class of genes that have recently gained significant attention. Despite lacking a clear evolutionary history, they are found in nearly all living organisms, from bacteria to humans, and they play important roles in diverse biological processes. The
[...] Read more.
Orphan Genes (OGs) are a mysterious class of genes that have recently gained significant attention. Despite lacking a clear evolutionary history, they are found in nearly all living organisms, from bacteria to humans, and they play important roles in diverse biological processes. The discovery of OGs was first made through comparative genomics followed by the identification of unique genes across different species. OGs tend to be more prevalent in species with larger genomes, such as plants and animals, and their evolutionary origins remain unclear but potentially arise from gene duplication, horizontal gene transfer (HGT), or de novo origination. Although their precise function is not well understood, OGs have been implicated in crucial biological processes such as development, metabolism, and stress responses. To better understand their significance, researchers are using a variety of approaches, including transcriptomics, functional genomics, and molecular biology. This review offers a comprehensive overview of the current knowledge of OGs in all domains of life, highlighting the possible role of dark transcriptomics in their evolution. More research is needed to fully comprehend the role of OGs in biology and their impact on various biological processes.
Full article
(This article belongs to the Special Issue 2022 Feature Papers by JDB’s Editorial Board Members)
►▼
Show Figures

Graphical abstract
Open AccessFeature PaperReview
An Emerging Animal Model for Querying the Role of Whole Genome Duplication in Development, Evolution, and Disease
J. Dev. Biol. 2023, 11(2), 26; https://doi.org/10.3390/jdb11020026 - 06 Jun 2023
Abstract
Whole genome duplication (WGD) or polyploidization can occur at the cellular, tissue, and organismal levels. At the cellular level, tetraploidization has been proposed as a driver of aneuploidy and genome instability and correlates strongly with cancer progression, metastasis, and the development of drug
[...] Read more.
Whole genome duplication (WGD) or polyploidization can occur at the cellular, tissue, and organismal levels. At the cellular level, tetraploidization has been proposed as a driver of aneuploidy and genome instability and correlates strongly with cancer progression, metastasis, and the development of drug resistance. WGD is also a key developmental strategy for regulating cell size, metabolism, and cellular function. In specific tissues, WGD is involved in normal development (e.g., organogenesis), tissue homeostasis, wound healing, and regeneration. At the organismal level, WGD propels evolutionary processes such as adaptation, speciation, and crop domestication. An essential strategy to further our understanding of the mechanisms promoting WGD and its effects is to compare isogenic strains that differ only in their ploidy. Caenorhabditis elegans (C. elegans) is emerging as an animal model for these comparisons, in part because relatively stable and fertile tetraploid strains can be produced rapidly from nearly any diploid strain. Here, we review the use of Caenorhabditis polyploids as tools to understand important developmental processes (e.g., sex determination, dosage compensation, and allometric relationships) and cellular processes (e.g., cell cycle regulation and chromosome dynamics during meiosis). We also discuss how the unique characteristics of the C. elegans WGD model will enable significant advances in our understanding of the mechanisms of polyploidization and its role in development and disease.
Full article
(This article belongs to the Special Issue Caenorhabditis elegans – a Model for Understanding Development and Disease)
►▼
Show Figures
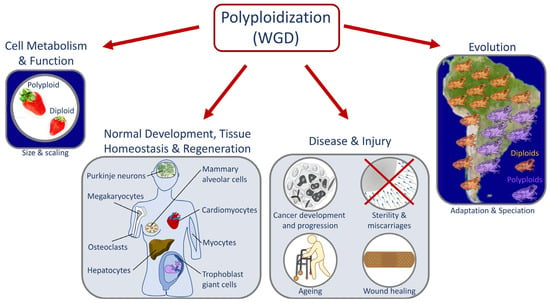
Figure 1
Open AccessFeature PaperReview
Evo Devo of the Vertebrates Integument
J. Dev. Biol. 2023, 11(2), 25; https://doi.org/10.3390/jdb11020025 - 05 Jun 2023
Abstract
All living jawed vertebrates possess teeth or did so ancestrally. Integumental surface also includes the cornea. Conversely, no other anatomical feature differentiates the clades so readily as skin appendages do, multicellular glands in amphibians, hair follicle/gland complexes in mammals, feathers in birds, and
[...] Read more.
All living jawed vertebrates possess teeth or did so ancestrally. Integumental surface also includes the cornea. Conversely, no other anatomical feature differentiates the clades so readily as skin appendages do, multicellular glands in amphibians, hair follicle/gland complexes in mammals, feathers in birds, and the different types of scales. Tooth-like scales are characteristic of chondrichthyans, while mineralized dermal scales are characteristic of bony fishes. Corneous epidermal scales might have appeared twice, in squamates, and on feet in avian lineages, but posteriorly to feathers. In contrast to the other skin appendages, the origin of multicellular glands of amphibians has never been addressed. In the seventies, pioneering dermal–epidermal recombination between chick, mouse and lizard embryos showed that: (1) the clade type of the appendage is determined by the epidermis; (2) their morphogenesis requires two groups of dermal messages, first for primordia formation, second for appendage final architecture; (3) the early messages were conserved during amniotes evolution. Molecular biology studies that have identified the involved pathways, extending those data to teeth and dermal scales, suggest that the different vertebrate skin appendages evolved in parallel from a shared placode/dermal cells unit, present in a common toothed ancestor, c.a. 420 mya.
Full article
(This article belongs to the Special Issue Development of the Skin in Vertebrates)
►▼
Show Figures
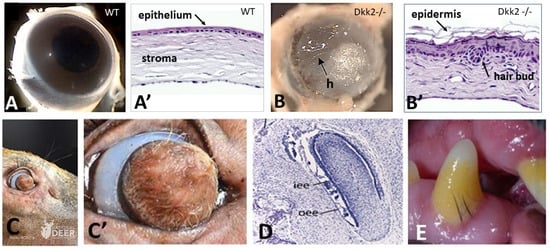
Figure 1
Open AccessFeature PaperArticle
Jak2 and Jaw Muscles Are Required for Buccopharyngeal Membrane Perforation during Mouth Development
J. Dev. Biol. 2023, 11(2), 24; https://doi.org/10.3390/jdb11020024 - 31 May 2023
Abstract
The mouth is a central feature of our face, without which we could not eat, breathe, or communicate. A critical and early event in mouth formation is the creation of a “hole” which connects the digestive system and the external environment. This hole,
[...] Read more.
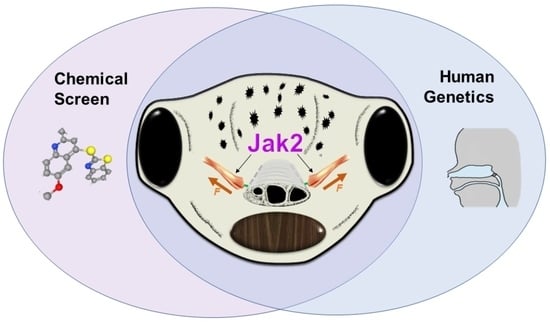
The mouth is a central feature of our face, without which we could not eat, breathe, or communicate. A critical and early event in mouth formation is the creation of a “hole” which connects the digestive system and the external environment. This hole, which has also been called the primary or embryonic mouth in vertebrates, is initially covered by a 1–2 cell layer thick structure called the buccopharyngeal membrane. When the buccopharyngeal membrane does not rupture, it impairs early mouth functions and may also lead to further craniofacial malformations. Using a chemical screen in an animal model (Xenopus laevis) and genetic data from humans, we determined that Janus kinase 2 (Jak2) has a role in buccopharyngeal membrane rupture. We have determined that decreased Jak2 function, using antisense morpholinos or a pharmacological antagonist, caused a persistent buccopharyngeal membrane as well as the loss of jaw muscles. Surprisingly, we observed that the jaw muscle compartments were connected to the oral epithelium that is continuous with the buccopharyngeal membrane. Severing such connections resulted in buccopharyngeal membrane buckling and persistence. We also noted puncta accumulation of F-actin, an indicator of tension, in the buccopharyngeal membrane during perforation. Taken together, the data has led us to a hypothesis that muscles are required to exert tension across the buccopharyngeal membrane, and such tension is necessary for its perforation.
Full article
(This article belongs to the Special Issue The 10th Anniversary of JDB: Feature Papers)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Transcription of HOX Genes Is Significantly Increased during Neuronal Differentiation of iPSCs Derived from Patients with Parkinson’s Disease
by
, , , , and
J. Dev. Biol. 2023, 11(2), 23; https://doi.org/10.3390/jdb11020023 - 25 May 2023
Abstract
Parkinson’s disease (PD) is the most serious movement disorder, but the actual cause of this disease is still unknown. Induced pluripotent stem cell-derived neural cultures from PD patients carry the potential for experimental modeling of underlying molecular events. We analyzed the RNA-seq data
[...] Read more.
Parkinson’s disease (PD) is the most serious movement disorder, but the actual cause of this disease is still unknown. Induced pluripotent stem cell-derived neural cultures from PD patients carry the potential for experimental modeling of underlying molecular events. We analyzed the RNA-seq data of iPSC-derived neural precursor cells (NPCs) and terminally differentiated neurons (TDNs) from healthy donors (HD) and PD patients with mutations in PARK2 published previously. The high level of transcription of HOX family protein-coding genes and lncRNA transcribed from the HOX clusters was revealed in the neural cultures from PD patients, while in HD NPCs and TDNs, the majority of these genes were not expressed or slightly transcribed. The results of this analysis were generally confirmed by qPCR. The HOX paralogs in the 3′ clusters were activated more strongly than the genes of the 5′ cluster. The abnormal activation of the HOX gene program upon neuronal differentiation in the cells of PD patients raises the possibility that the abnormal expression of these key regulators of neuronal development impacts PD pathology. Further research is needed to investigate this hypothesis.
Full article
(This article belongs to the Special Issue 2022 Feature Papers by JDB’s Editorial Board Members)
►▼
Show Figures
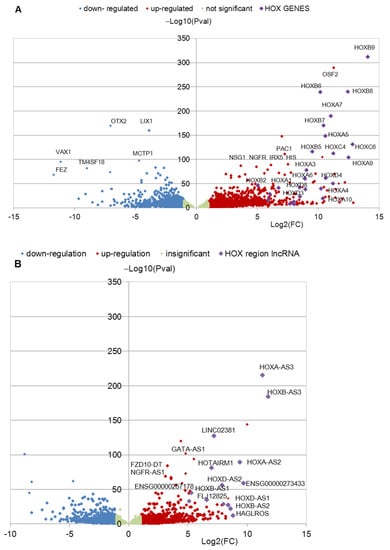
Figure 1
Open AccessFeature PaperArticle
Osteoderm Development during the Regeneration Process in Eurylepis taeniolata Blyth, 1854 (Scincidae, Sauria, Squamata)
J. Dev. Biol. 2023, 11(2), 22; https://doi.org/10.3390/jdb11020022 - 24 May 2023
Abstract
Osteoderms are bony structures that develop within the dermal layer of the skin in vertebrates and are very often found in different lizard families. Lizard osteoderms are diverse in topography, morphology, and microstructure. Of particular interest are the compound osteoderms of skinks, which
[...] Read more.
Osteoderms are bony structures that develop within the dermal layer of the skin in vertebrates and are very often found in different lizard families. Lizard osteoderms are diverse in topography, morphology, and microstructure. Of particular interest are the compound osteoderms of skinks, which are a complex of several bone elements known as osteodermites. We present new data on the development and regeneration of compound osteoderms based on the results of a histological and Computed Microtomography (micro-CT) study of a scincid lizard: Eurylepis taeniolata. The specimens studied are stored in the herpetological collections of the Saint-Petersburg State University and Zoological Institute of the Russian Academy of Sciences located in St. Petersburg, Russia. The topography of osteoderms in the integuments of the original tail area and its regenerated part was studied. A comparative histological description of the original and regenerated osteoderms of Eurylepis taeniolata is presented for the first time. The first description of the development of compound osteoderm microstructure in the process of caudal regeneration is also presented.
Full article
(This article belongs to the Special Issue Development of the Skin in Vertebrates)
►▼
Show Figures
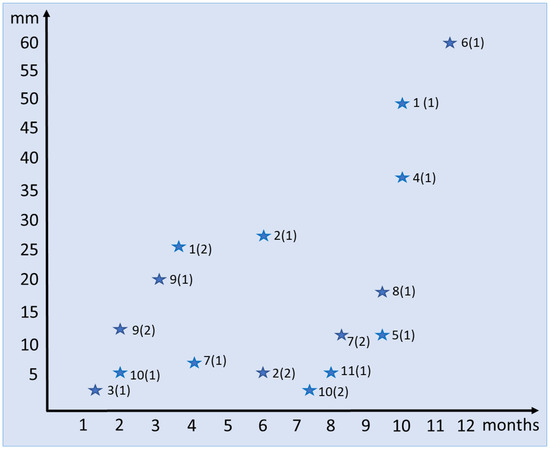
Figure 1
Open AccessFeature PaperReview
Genetic and Epigenetic Regulation of Drosophila Oocyte Determination
J. Dev. Biol. 2023, 11(2), 21; https://doi.org/10.3390/jdb11020021 - 24 May 2023
Abstract
Primary oocyte determination occurs in many organisms within a germ line cyst, a multicellular structure composed of interconnected germ cells. However, the structure of the cyst is itself highly diverse, which raises intriguing questions about the benefits of this stereotypical multicellular environment for
[...] Read more.
Primary oocyte determination occurs in many organisms within a germ line cyst, a multicellular structure composed of interconnected germ cells. However, the structure of the cyst is itself highly diverse, which raises intriguing questions about the benefits of this stereotypical multicellular environment for female gametogenesis. Drosophila melanogaster is a well-studied model for female gametogenesis, and numerous genes and pathways critical for the determination and differentiation of a viable female gamete have been identified. This review provides an up-to-date overview of Drosophila oocyte determination, with a particular emphasis on the mechanisms that regulate germ line gene expression.
Full article
(This article belongs to the Special Issue The 10th Anniversary of JDB: Feature Papers)
►▼
Show Figures
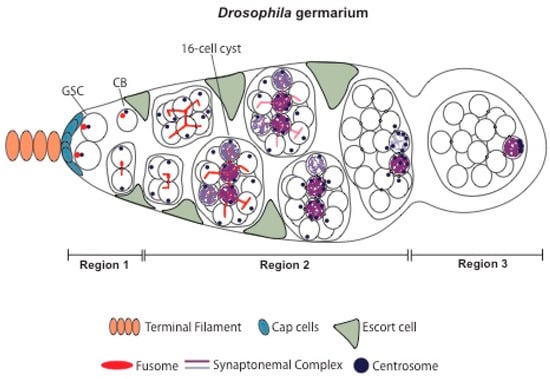
Figure 1
Open AccessFeature PaperArticle
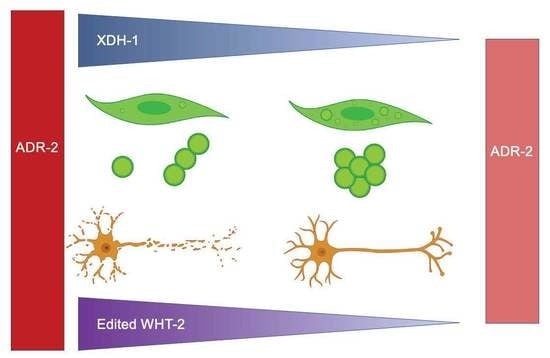
Attenuation of Dopaminergic Neurodegeneration in a C. elegans Parkinson’s Model through Regulation of Xanthine Dehydrogenase (XDH-1) Expression by the RNA Editase, ADR-2
by
, , , , , and
J. Dev. Biol. 2023, 11(2), 20; https://doi.org/10.3390/jdb11020020 - 22 May 2023
Abstract
Differential RNA editing by adenosine deaminases that act on RNA (ADARs) has been implicated in several neurological disorders, including Parkinson’s disease (PD). Here, we report results of a RNAi screen of genes differentially regulated in adr-2 mutants, normally encoding the only catalytically active
[...] Read more.
Differential RNA editing by adenosine deaminases that act on RNA (ADARs) has been implicated in several neurological disorders, including Parkinson’s disease (PD). Here, we report results of a RNAi screen of genes differentially regulated in adr-2 mutants, normally encoding the only catalytically active ADAR in Caenorhabditis elegans, ADR-2. Subsequent analysis of candidate genes that alter the misfolding of human α-synuclein (α-syn) and dopaminergic neurodegeneration, two PD pathologies, reveal that reduced expression of xdh-1, the ortholog of human xanthine dehydrogenase (XDH), is protective against α-synuclein-induced dopaminergic neurodegeneration. Further, RNAi experiments show that WHT-2, the worm ortholog of the human ABCG2 transporter and a predicted interactor of XDH-1, is the rate-limiting factor in the ADR-2, XDH-1, WHT-2 system for dopaminergic neuroprotection. In silico structural modeling of WHT-2 indicates that the editing of one nucleotide in the wht-2 mRNA leads to the substitution of threonine with alanine at residue 124 in the WHT-2 protein, changing hydrogen bonds in this region. Thus, we propose a model where wht-2 is edited by ADR-2, which promotes optimal export of uric acid, a known substrate of WHT-2 and a product of XDH-1 activity. In the absence of editing, uric acid export is limited, provoking a reduction in xdh-1 transcription to limit uric acid production and maintain cellular homeostasis. As a result, elevation of uric acid is protective against dopaminergic neuronal cell death. In turn, increased levels of uric acid are associated with a decrease in ROS production. Further, downregulation of xdh-1 is protective against PD pathologies because decreased levels of XDH-1 correlate to a concomitant reduction in xanthine oxidase (XO), the form of the protein whose by-product is superoxide anion. These data indicate that modifying specific targets of RNA editing may represent a promising therapeutic strategy for PD.
Full article
(This article belongs to the Special Issue Caenorhabditis elegans – a Model for Understanding Development and Disease)
►▼
Show Figures
Graphical abstract
Open AccessArticle
The Presence of Two MyoD Genes in a Subset of Acanthopterygii Fish Is Associated with a Polyserine Insert in MyoD1
by
, , , and
J. Dev. Biol. 2023, 11(2), 19; https://doi.org/10.3390/jdb11020019 - 28 Apr 2023
Abstract
The MyoD gene was duplicated during the teleost whole genome duplication and, while a second MyoD gene (MyoD2) was subsequently lost from the genomes of some lineages (including zebrafish), many fish lineages (including Alcolapia species) have retained both MyoD paralogues. Here
[...] Read more.
The MyoD gene was duplicated during the teleost whole genome duplication and, while a second MyoD gene (MyoD2) was subsequently lost from the genomes of some lineages (including zebrafish), many fish lineages (including Alcolapia species) have retained both MyoD paralogues. Here we reveal the expression patterns of the two MyoD genes in Oreochromis (Alcolapia) alcalica using in situ hybridisation. We report our analysis of MyoD1 and MyoD2 protein sequences from 54 teleost species, and show that O. alcalica, along with some other teleosts, include a polyserine repeat between the amino terminal transactivation domains (TAD) and the cysteine-histidine rich region (H/C) in MyoD1. The evolutionary history of MyoD1 and MyoD2 is compared to the presence of this polyserine region using phylogenetics, and its functional relevance is tested using overexpression in a heterologous system to investigate subcellular localisation, stability, and activity of MyoD proteins that include and do not include the polyserine region.
Full article
(This article belongs to the Special Issue The 10th Anniversary of JDB: Feature Papers)
►▼
Show Figures
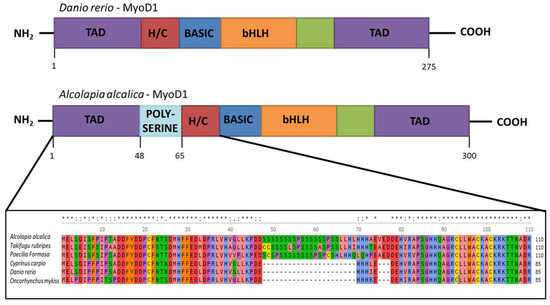
Figure 1
Open AccessFeature PaperArticle
Reproductive-Toxicity-Related Endpoints in C. elegans Are Consistent with Reduced Concern for Dimethylarsinic Acid Exposure Relative to Inorganic Arsenic
J. Dev. Biol. 2023, 11(2), 18; https://doi.org/10.3390/jdb11020018 - 26 Apr 2023
Abstract
Exposures to arsenic and mercury are known to pose significant threats to human health; however, the effects specific to organic vs. inorganic forms are not fully understood. Caenorhabditis elegans’ (C. elegans) transparent cuticle, along with the conservation of key genetic pathways
[...] Read more.
Exposures to arsenic and mercury are known to pose significant threats to human health; however, the effects specific to organic vs. inorganic forms are not fully understood. Caenorhabditis elegans’ (C. elegans) transparent cuticle, along with the conservation of key genetic pathways regulating developmental and reproductive toxicology (DART)-related processes such as germ stem cell renewal and differentiation, meiosis, and embryonic tissue differentiation and growth, support this model’s potential to address the need for quicker and more dependable testing methods for DART hazard identification. Organic and inorganic forms of mercury and arsenic had different effects on reproductive-related endpoints in C. elegans, with methylmercury (meHgCl) having effects at lower concentrations than mercury chloride (HgCl2), and sodium arsenite (NaAsO2) having effects at lower concentrations than dimethylarsinic acid (DMA). Progeny to adult ratio changes and germline apoptosis were seen at concentrations that also affected gravid adult gross morphology. For both forms of arsenic tested, germline histone regulation was altered at concentrations below those that affected progeny/adult ratios, while concentrations for these two endpoints were similar for the mercury compounds. These C. elegans findings are consistent with corresponding mammalian data, where available, suggesting that small animal model test systems may help to fill critical data gaps by contributing to weight of evidence assessments.
Full article
(This article belongs to the Special Issue Caenorhabditis elegans – a Model for Understanding Development and Disease)
►▼
Show Figures
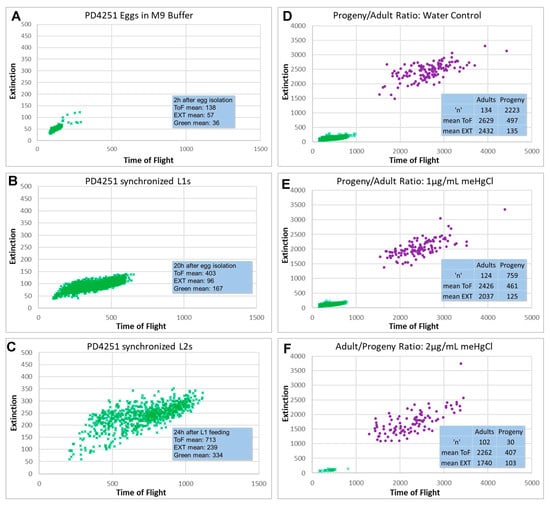
Figure 1
Open AccessArticle
Selecting Normalizers for MicroRNA RT-qPCR Expression Analysis in Murine Preimplantation Embryos and the Associated Conditioned Culture Media
J. Dev. Biol. 2023, 11(2), 17; https://doi.org/10.3390/jdb11020017 - 04 Apr 2023
Abstract
►▼
Show Figures
Normalizing RT-qPCR miRNA datasets that encompass numerous preimplantation embryo stages requires the identification of miRNAs that may be used as stable reference genes. A need has also arisen for the normalization of the accompanying conditioned culture media as extracellular miRNAs may serve as
[...] Read more.
Normalizing RT-qPCR miRNA datasets that encompass numerous preimplantation embryo stages requires the identification of miRNAs that may be used as stable reference genes. A need has also arisen for the normalization of the accompanying conditioned culture media as extracellular miRNAs may serve as biomarkers of embryo developmental competence. Here, we evaluate the stability of six commonly used miRNA normalization candidates, as well as small nuclear U6, using five different means of evaluation (BestKeeper, NormFinder, geNorm, the comparative Delta Ct method and RefFinder comprehensive analysis) to assess their stability throughout murine preimplantation embryo development from the oocyte to the late blastocyst stages, both in whole embryos and the associated conditioned culture media. In descending order of effectiveness, miR-16, miR-191 and miR-106 were identified as the most stable individual reference miRNAs for developing whole CD1 murine preimplantation embryos, while miR-16, miR-106 and miR-103 were ideal for the conditioned culture media. Notably, the widely used U6 reference was among the least appropriate for normalizing both whole embryo and conditioned media miRNA datasets. Incorporating multiple reference miRNAs into the normalization basis via a geometric mean was deemed beneficial, and combinations of each set of stable miRNAs are further recommended, pending validation on a per experiment basis.
Full article

Figure 1
Open AccessCommunication
Comparison of Pronase versus Manual Dechorionation of Zebrafish Embryos for Small Molecule Treatments
J. Dev. Biol. 2023, 11(2), 16; https://doi.org/10.3390/jdb11020016 - 28 Mar 2023
Abstract
Zebrafish are a powerful animal model for small molecule screening. Small molecule treatments of zebrafish embryos usually require that the chorion, an acellular envelope enclosing the embryo, is removed in order for chemical compounds to access the embryo from the bath medium. For
[...] Read more.
Zebrafish are a powerful animal model for small molecule screening. Small molecule treatments of zebrafish embryos usually require that the chorion, an acellular envelope enclosing the embryo, is removed in order for chemical compounds to access the embryo from the bath medium. For large-scale studies requiring hundreds of embryos, manual dechorionation, using forceps, can be a time-consuming and limiting process. Pronase is a non-specific protease that is widely used as an enzymatic alternative for dechorionating zebrafish embryos. However, whether pronase treatments alter the effects of subsequent small molecule treatments has not been addressed. Here, we provide a detailed protocol for large-scale pronase dechorionation of zebrafish embryos. We tested whether pronase treatment can influence the efficacy of drug treatments in zebrafish embryos. We used a zebrafish model for Duchenne muscular dystrophy (DMD) to investigate whether the efficacies of trichostatin-A (TSA) or salermide + oxamflatin, small molecule inhibitors known to ameliorate the zebrafish dmd muscle degeneration phenotype, are significantly altered when embryos are treated with pronase versus manual dechorionation. We also tested the effects of pronase on the ability of the anthracycline cancer drug doxorubicin to induce cardiotoxicity in zebrafish embryos. When comparing pronase- versus forceps-dechorionated embryos used in these small molecule treatments, we found no appreciable effects of pronase on animal survival or on the effects of the small molecules. The significant difference that was detected was a small improvement in the ability of salermide + oxamflatin to ameliorate the dmd phenotype in pronase-treated embryos when compared with manual dechorionation. Our study supports the use of pronase treatment as a dechorionation method for zebrafish drug screening experiments.
Full article
(This article belongs to the Special Issue Zebrafish—a Model System for Developmental Biology II)
►▼
Show Figures
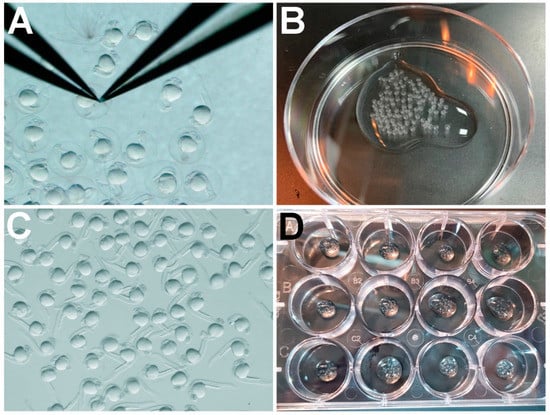

Figure 1
Open AccessFeature PaperArticle
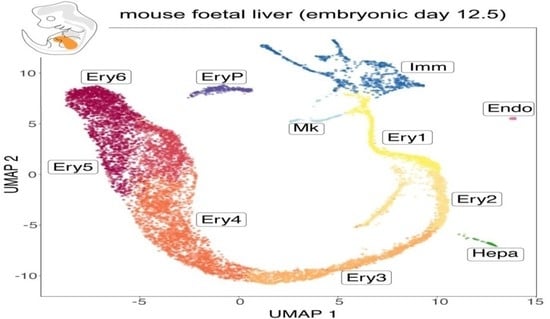
A Refined Single Cell Landscape of Haematopoiesis in the Mouse Foetal Liver
by
, , , , and
J. Dev. Biol. 2023, 11(2), 15; https://doi.org/10.3390/jdb11020015 - 23 Mar 2023
Abstract
During prenatal life, the foetal liver is colonised by several waves of haematopoietic progenitors to act as the main haematopoietic organ. Single cell (sc) RNA-seq has been used to identify foetal liver cell types via their transcriptomic signature and to compare gene expression
[...] Read more.
During prenatal life, the foetal liver is colonised by several waves of haematopoietic progenitors to act as the main haematopoietic organ. Single cell (sc) RNA-seq has been used to identify foetal liver cell types via their transcriptomic signature and to compare gene expression patterns as haematopoietic development proceeds. To obtain a refined single cell landscape of haematopoiesis in the foetal liver, we have generated a scRNA-seq dataset from a whole mouse E12.5 liver that includes a larger number of cells than prior datasets at this stage and was obtained without cell type preselection to include all liver cell populations. We combined mining of this dataset with that of previously published datasets at other developmental stages to follow transcriptional dynamics as well as the cell cycle state of developing haematopoietic lineages. Our findings corroborate several prior reports on the timing of liver colonisation by haematopoietic progenitors and the emergence of differentiated lineages and provide further molecular characterisation of each cell population. Extending these findings, we demonstrate the existence of a foetal intermediate haemoglobin profile in the mouse, similar to that previously identified in humans, and a previously unidentified population of primitive erythroid cells in the foetal liver.
Full article
(This article belongs to the Topic Advances in Red Blood Cells Research)
►▼
Show Figures
Graphical abstract
Open AccessReview
Principles of Zebrafish Nephron Segment Development
J. Dev. Biol. 2023, 11(1), 14; https://doi.org/10.3390/jdb11010014 - 18 Mar 2023
Cited by 2
Abstract
Nephrons are the functional units which comprise the kidney. Each nephron contains a number of physiologically unique populations of specialized epithelial cells that are organized into discrete domains known as segments. The principles of nephron segment development have been the subject of many
[...] Read more.
Nephrons are the functional units which comprise the kidney. Each nephron contains a number of physiologically unique populations of specialized epithelial cells that are organized into discrete domains known as segments. The principles of nephron segment development have been the subject of many studies in recent years. Understanding the mechanisms of nephrogenesis has enormous potential to expand our knowledge about the basis of congenital anomalies of the kidney and urinary tract (CAKUT), and to contribute to ongoing regenerative medicine efforts aimed at identifying renal repair mechanisms and generating replacement kidney tissue. The study of the zebrafish embryonic kidney, or pronephros, provides many opportunities to identify the genes and signaling pathways that control nephron segment development. Here, we describe recent advances of nephron segment patterning and differentiation in the zebrafish, with a focus on distal segment formation.
Full article
(This article belongs to the Special Issue Zebrafish—a Model System for Developmental Biology II)
►▼
Show Figures

Figure 1

Journal Menu
► ▼ Journal Menu-
- JDB Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Topical Collections
- Article Processing Charge
- Indexing & Archiving
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Conferences
- Editorial Office
- 10th Anniversary of JDB
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
BioChem, Biology, Biomolecules, JDB, Metabolites
"Metabolites" as Metabolomics for Neuropsychiatric Diseases
Topic Editors: Reiji Yoshimura, Hiroshi Kunugi, Takahiro Kato, Naomichi OkamotoDeadline: 25 November 2023

Conferences
Special Issues
Special Issue in
JDB
Recent Advances in Skeletal Development and Diseases
Guest Editor: Tao YangDeadline: 31 August 2023
Special Issue in
JDB
10th Anniversary of JDB—Recent Advances in Wnt Signaling in Development and Regeneration
Guest Editor: Ryan RangeDeadline: 30 October 2023
Special Issue in
JDB
Caenorhabditis elegans – a Model for Understanding Development and Disease
Guest Editors: Ann K. Corsi, Andy GoldenDeadline: 20 November 2023
Special Issue in
JDB
The 10th Anniversary of JDB: Feature Papers
Guest Editor: Simon J. ConwayDeadline: 31 December 2023
Topical Collections
Topical Collection in
JDB
Drosophila - A Model System for Developmental Biology
Collection Editor: Nicholas Tolwinski
Topical Collection in
JDB
Hedgehog Signaling in Embryogenesis
Collection Editors: Henk Roelink, Kay Grobe