Journal Description
Biophysica
Biophysica
is an international, peer-reviewed, open access journal on applying the methods of physics, chemistry, and math to study biological systems, published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 19.9 days after submission; acceptance to publication is undertaken in 4.9 days (median values for papers published in this journal in the first half of 2023).
- Recognition of reviewers: APC discount vouchers, optional signed peer review and reviewer names are published annually in the journal.
- Biophysica is a companion journal of IJMS.
Latest Articles
The Dynamic Behavior of a Single Semiflexible Ring Chain in a Linear Polymer Matrix
Biophysica 2023, 3(3), 476-484; https://doi.org/10.3390/biophysica3030031 - 26 Jul 2023
Abstract
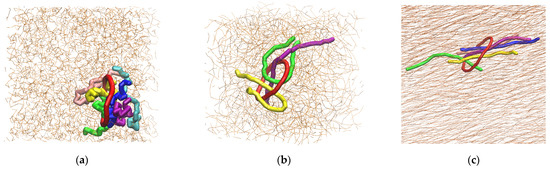
We studied the dynamic behavior of a single semiflexible ring in linear chain matrix based on a coarse-grained model using the molecular dynamics simulation approach. We found that that ring chains’ hollow centers are frequently filled with linear chains. However, as the rigidity
[...] Read more.
We studied the dynamic behavior of a single semiflexible ring in linear chain matrix based on a coarse-grained model using the molecular dynamics simulation approach. We found that that ring chains’ hollow centers are frequently filled with linear chains. However, as the rigidity of the linear chains increases, the linear chains arranged parallel to each other and the ring chain are temporary caged. As a result, the swing movement in the normal direction of the ring is significantly limited, and the relaxation time in the normal direction increases significantly. Our findings can help to understand the physical mechanism of the movement of the ring chain in ring–linear polymer blends at the microscopic level.
Full article
(This article belongs to the Special Issue Molecular Structure and Simulation in Biological System)
►
Show Figures
Open AccessArticle
Exploring the Dynamics of Holo-Shikimate Kinase through Molecular Mechanics
Biophysica 2023, 3(3), 463-475; https://doi.org/10.3390/biophysica3030030 - 21 Jul 2023
Abstract
Understanding the connection between local and global dynamics can provide valuable insights into enzymatic function and may contribute to the development of novel strategies for enzyme modulation. In this work, we investigated the dynamics at both the global and local (active site) levels
[...] Read more.
Understanding the connection between local and global dynamics can provide valuable insights into enzymatic function and may contribute to the development of novel strategies for enzyme modulation. In this work, we investigated the dynamics at both the global and local (active site) levels of Shikimate Kinase (SK) through microsecond time-scale molecular dynamics (MD) simulations of the holoenzyme in the product state. Our focus was on the wild-type (WT) enzyme and two mutants (R116A and R116K) which are known for their reduced catalytic activity. Through exploring the dynamics of these variants, we gained insights into the role of residue R116 and its contribution to overall SK dynamics. We argue that the connection between local and global dynamics can be attributed to local frustration near the mutated residue which perturbs the global protein dynamics.
Full article
(This article belongs to the Collection Feature Papers in Biophysics)
►▼
Show Figures
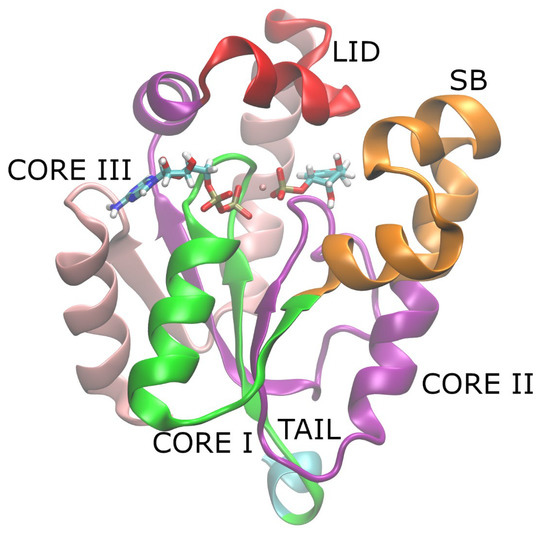
Figure 1
Open AccessArticle
A Structure-Guided Designed Small Molecule Is an Anticancer Agent and Inhibits the Apoptosis-Related MCL-1 Protein
by
, , , , , , , , , , and
Biophysica 2023, 3(3), 446-462; https://doi.org/10.3390/biophysica3030029 - 07 Jul 2023
Abstract
Cancer resistance to chemotherapy and radiation therapies presents significant challenges, necessitating the exploration of alternative approaches. Targeting specific proteins at the molecular level, particularly their active sites, holds promise in addressing this issue. We investigated the potential of 4′-methoxy-2-nitrochalcone (MNC) as an MCL-1
[...] Read more.
Cancer resistance to chemotherapy and radiation therapies presents significant challenges, necessitating the exploration of alternative approaches. Targeting specific proteins at the molecular level, particularly their active sites, holds promise in addressing this issue. We investigated the potential of 4′-methoxy-2-nitrochalcone (MNC) as an MCL-1 inhibitor, examining its chemical and structural characteristics to elucidate its biological activity and guide the selection of potential candidates. We conducted a docking study, followed by synthesis, structural characterization, theoretical calculations, and in vitro experiments to comprehensively evaluate MNC. The docking results revealed MNC’s excellent binding within the active site of MCL-1. At 50 µM, MNC demonstrated 99% inhibition of HCT116 cell proliferation, with an IC50 value of 15.18 µM after 24 h. Treatment with MNC at 30.36 and 15.18 µM resulted in reduced cell density. Notably, MNC exhibited marked cytotoxicity at concentrations of 15.58 µM and 7.79 µM, inducing high frequencies of plasma membrane rupture and apoptosis, respectively. Our findings highlight the significant biological potential of MNC as an MCL-1 inhibitor. Furthermore, we propose exploring chalcones with hydrogen bond acceptor substituents as promising candidates for studying inhibitors targeting this protein. In conclusion, our study addresses the challenge of cancer resistance by investigating MNC as an MCL-1 inhibitor. Through detailed characterization and experimental validation, we establish the efficacof MNC in inhibiting cell proliferation and inducing cytotoxic effects. These results underscore the potential of MNC as a valuable therapeutic agent and suggest the use of chalcones with hydrogen bond acceptor substituents as a basis for developing novel MCL-1 inhibitors.
Full article
(This article belongs to the Collection Feature Papers in Biophysics)
►▼
Show Figures

Figure 1
Open AccessArticle
Screening for Bioactive Metabolites in Leaves, Branches, and Roots of Mansoa hirsuta: Phytochemical, Toxicological and Antioxidant Aspects
by
, , , , , , , , , , and
Biophysica 2023, 3(3), 425-445; https://doi.org/10.3390/biophysica3030028 - 28 Jun 2023
Abstract
In this study, secondary metabolites, toxicology and antioxidant properties of chloroform fractions from leaves (FCFMh), branches (FCGMh), and roots (FCRMh) of Mansoa hirsuta were investigated. The phytochemical screening detected flavonoids, especially chalcones. Through Liquid chromatography with mass spectrometry—LC–MS analysis, the flavonoids (isoorientin-2″-O
[...] Read more.
In this study, secondary metabolites, toxicology and antioxidant properties of chloroform fractions from leaves (FCFMh), branches (FCGMh), and roots (FCRMh) of Mansoa hirsuta were investigated. The phytochemical screening detected flavonoids, especially chalcones. Through Liquid chromatography with mass spectrometry—LC–MS analysis, the flavonoids (isoorientin-2″-O-arabinoside), triterpenes (oleanolic acid and ursolic acid) and ceramide (phytosphingosine) were identified. From the Artemia salina assay, the fraction FCGMh was the most toxic (LC50 = 64.21 µg·mL−1), followed by FCRMh (LC50 = 87.61 µg·mL−1) and FCFMh (LC50 = 421.9 µg·mL−1). Concerning the cytotoxic potential, the root fraction (IC50 16.48 μg mL−1) displayed the highest cytotoxicity against the breast cancer cell line (4T1), followed by leaves (IC50 33.13 μg mL−1) and branches (IC50 of 47.13 μg mL−1). In conclusion, all the fractions of M. hirsuta showed cytotoxicity at the highest concentrations; however, remarkable biological properties were found for the root fractions. Computational analysis was performed using a molecular docking and pharmacophore approach to understand the antioxidant activity of its major metabolites.
Full article
(This article belongs to the Special Issue Molecular Structure and Simulation in Biological System)
►▼
Show Figures
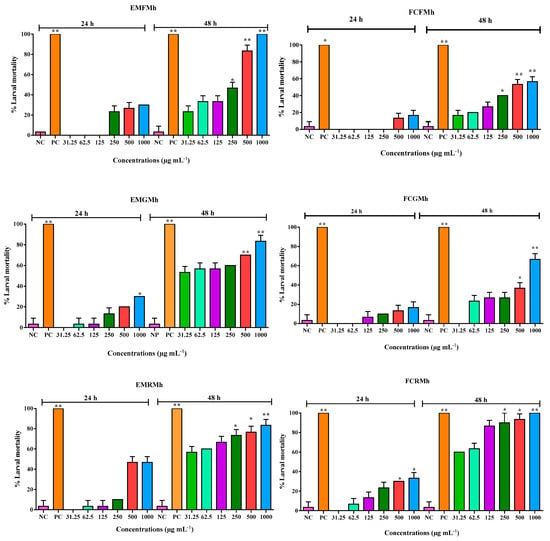
Figure 1
Open AccessArticle
Adsorption of Heparin-Binding Fragments of Fibronectin onto Hydrophobic Surfaces
Biophysica 2023, 3(3), 409-424; https://doi.org/10.3390/biophysica3030027 - 23 Jun 2023
Abstract
Fibronectin is a multi-domain, extracellular matrix protein that plays a number of biological roles. As the adsorption of fibronectin onto the surface of implanted devices can lead to an inflammatory response or bacterial colonisation, understanding the interaction of fibronectin with material surfaces is
[...] Read more.
Fibronectin is a multi-domain, extracellular matrix protein that plays a number of biological roles. As the adsorption of fibronectin onto the surface of implanted devices can lead to an inflammatory response or bacterial colonisation, understanding the interaction of fibronectin with material surfaces is important in the design of materials for biomedical applications. This, however, relies on having knowledge of the molecular-scale behaviour of proteins, which is difficult to investigate experimentally. In this paper, we used molecular dynamics simulations to investigate the adsorption of heparin-binding fibronectin domains onto hydrophobic surfaces. Despite the high similarity between these, their adsorption differs both in terms of the strength and the specificity of this, indicating that relatively small changes in protein structure can lead to significant changes in adsorption behaviour. This suggests that the interplay between protein structure and surface chemistry is vital for understanding the protein adsorption process and the design of novel biomaterials.
Full article
(This article belongs to the Special Issue Protein Engineering: The Present and the Future 2.0)
►▼
Show Figures
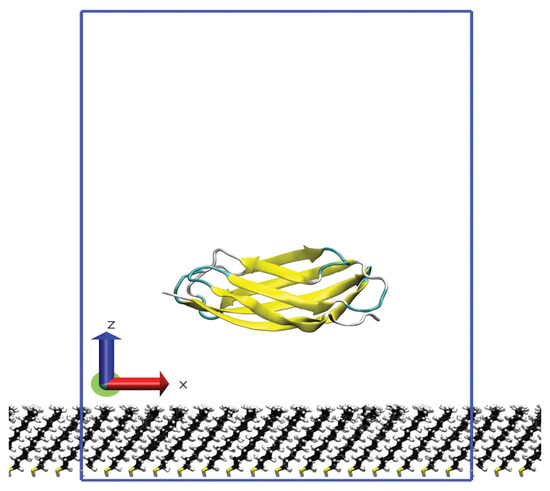
Figure 1
Open AccessArticle
A Two-Species Finite Volume Scalar Model for Modeling the Diffusion of Poly(lactic-co-glycolic acid) into a Coronary Arterial Wall from a Single Half-Embedded Drug Eluting Stent Strut
Biophysica 2023, 3(2), 385-408; https://doi.org/10.3390/biophysica3020026 - 15 Jun 2023
Abstract
This paper outlines the methodology and results for a two-species finite volume scalar computational drug transport model developed for simulating the mass transport of Poly(lactic-co-glycolic acid (PLGA)) from a half-embedded single strut implanted in a coronary arterial vessel wall. The mathematical drug transport
[...] Read more.
This paper outlines the methodology and results for a two-species finite volume scalar computational drug transport model developed for simulating the mass transport of Poly(lactic-co-glycolic acid (PLGA)) from a half-embedded single strut implanted in a coronary arterial vessel wall. The mathematical drug transport model incorporates the convection-diffusion equation in scalar form (dimensionless) with a two-species (free-drug and bound-drug) mass transport setup, including reversible equilibrium reaction source terms for the free and bound-drug states to account for the pharmaco-kinetic reactions in the arterial wall. The relative reaction rates of the added source terms control the interconversion of the drug between the free and bound-drug states. The model is solved by a 2D finite-volume method for discretizing and solving the free and bound drug transport equations with anisotropic vascular drug diffusivities. This model is an improvement over previously developed models using the finite-difference and finite element method. A dimensionless characteristic scaling pre-analysis was conducted a priori to evaluate the significance of implementing the reaction source terms in the transport equations. This paper reports the findings of an investigation of the interstitial flow profile into the arterial wall and the free and bound drug diffusion profiles with a parametric study of varying the polymer drug concentration (low and high), tortuosity, porosity, and Peclet and DamKöhler numbers over the course of 400 h (16.67 days). The results also reveal how a single species drug delivery model that neglects both a reversible binding reaction source term and the porosity and tortuosity of the arterial wall cannot accurately predict the distribution of both the free and bound drug.
Full article
(This article belongs to the Special Issue Molecular Structure and Simulation in Biological System)
►▼
Show Figures
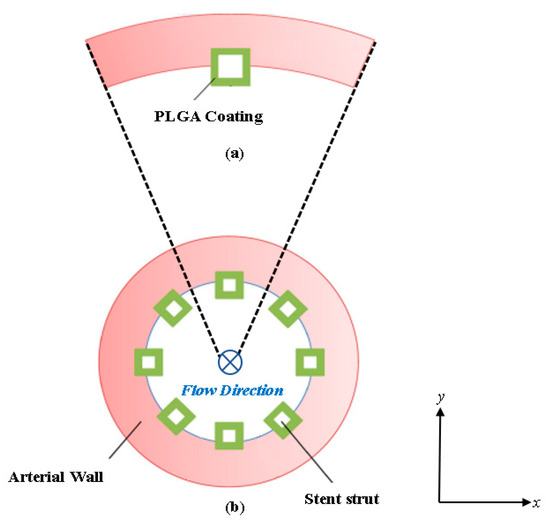
Figure 1
Open AccessArticle
Computational Modeling of the Neurofibromin-Stimulated Guanosine Triphosphate Hydrolysis by the KRas Protein
by
and
Biophysica 2023, 3(2), 373-384; https://doi.org/10.3390/biophysica3020025 - 31 May 2023
Abstract
We report the results of computational studies of the guanosine triphosphate (GTP) hydrolysis in the active site of the KRas-NF1 protein complex, where KRas stands for the K-isoform of the Ras (ras sarcoma) protein and NF1 (neurofbromin-1) is the activating protein. The model
[...] Read more.
We report the results of computational studies of the guanosine triphosphate (GTP) hydrolysis in the active site of the KRas-NF1 protein complex, where KRas stands for the K-isoform of the Ras (ras sarcoma) protein and NF1 (neurofbromin-1) is the activating protein. The model system was constructed using coordinates of heavy atoms from the crystal structure PDB ID 6OB2 with the GTP analog GMPPNP. Large-scale classical molecular dynamics (MD) calculations were performed to analyze conformations of the enzyme-substrate complexes. The Gibbs energy profiles for the hydrolysis reaction were computed using MD simulations with quantum mechanics/molecular mechanics (QM/MM) interaction potentials. The density functional theory DFT(ωB97X-D3/6-31G**) approach was applied in QM and the CHARMM36 force field parameters in MM. The most likely scenario of the chemical step of the GTP hydrolysis in KRas-NF1 corresponds to the water-assisted mechanism of the formation of the inorganic phosphate coupled with the dissociation of GTP to GDP.
Full article
(This article belongs to the Special Issue Molecular Structure and Simulation in Biological System)
►▼
Show Figures
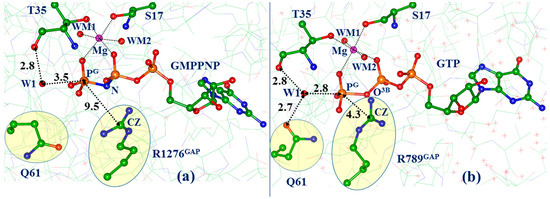
Figure 1
Open AccessArticle
Decomposition of Small Molecules for Fragment-Based Drug Design
Biophysica 2023, 3(2), 362-372; https://doi.org/10.3390/biophysica3020024 - 24 May 2023
Abstract
In the drug design process, a frequent task is the decomposition of small molecules into fragments. There exist a number of approaches and methods to break molecules into fragments. However, a method that allows the decomposition of molecules into non-overlapping fragments that is
[...] Read more.
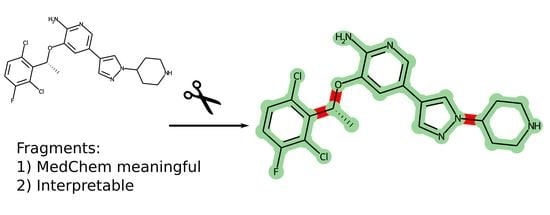
In the drug design process, a frequent task is the decomposition of small molecules into fragments. There exist a number of approaches and methods to break molecules into fragments. However, a method that allows the decomposition of molecules into non-overlapping fragments that is meaningful in terms of medicinal chemistry is absent, and in this work, we present a new simple approach for the decomposition of molecules—MedChemFrag. It aims to break drug-like molecules into a set of rings and linkers, which are close to the perception of “fragments” by medicinal chemists. In contrast to most previous efforts aimed at breaking molecules using retrosynthetic feasible rules, our approach strives to preserve the functional groups, which may reveal the specific interaction pattern, e.g., the amide groups.
Full article
(This article belongs to the Special Issue Molecular Structure and Simulation in Biological System)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Phenotypic and Biomechanical Characteristics of Human Fetal Neural Progenitor Cells Exposed to Pesticide Compounds
by
, , , and
Biophysica 2023, 3(2), 348-361; https://doi.org/10.3390/biophysica3020023 - 18 May 2023
Abstract
Various forms of pesticides have been reported to be among the environmental toxicants, which are detrimental to human health. The active ingredients of these formulations can enter the human body through air, food, or water. Epidemiological studies suggest that these compounds strongly affect
[...] Read more.
Various forms of pesticides have been reported to be among the environmental toxicants, which are detrimental to human health. The active ingredients of these formulations can enter the human body through air, food, or water. Epidemiological studies suggest that these compounds strongly affect the developing brain in fetal and infant stages due to their ability to breach the underdeveloped blood–brain barrier. Since neural progenitor stem cells (NPCs) in the developing brain are the most vulnerable to these compounds, the mechanisms by which NPCs experience toxicity upon exposure to these chemicals must be investigated. Here, we assessed the viability of human fetal NPCs in 2D cultures in the presence of the active ingredients of six widely used pesticides using Live/Dead® and Hoechst staining. The IC50 values ranged from 4.1–201 μM. A significant drop in cell viability with increasing toxicant concentration (p < 0.01) was noted, with the order of toxicity being malathion < 4-aminopyridine < methoprene < prallethrin < temephos < pyriproxyfen. Changes in cellular biomechanical characteristics (Young’s modulus, tether force, membrane tension, and tether radius) were quantified using atomic force microscopy, whereas cell migration was elucidated over 48 h using a customized wound-healing assay. The Young’s modulus of fetal NPCs exposed to IC50/2 doses of these compounds was reduced by 38–70% and that of those exposed to IC50 doses was reduced by 71–80% (p < 0.001 vs. controls for both; p < 0.01 for IC50 vs. IC50/2 for each compound). Similar patterns were noted for tether forces and membrane tension in fetal NPCs. NPC migration was found to be compound type- and dose-dependent. These results attest to the significant detrimental effects of these compounds on various aspects of the human fetal NPC phenotype, and the utility of cell mechanics as a marker to assess developmental neurotoxicity.
Full article
(This article belongs to the Collection Feature Papers in Biophysics)
►▼
Show Figures
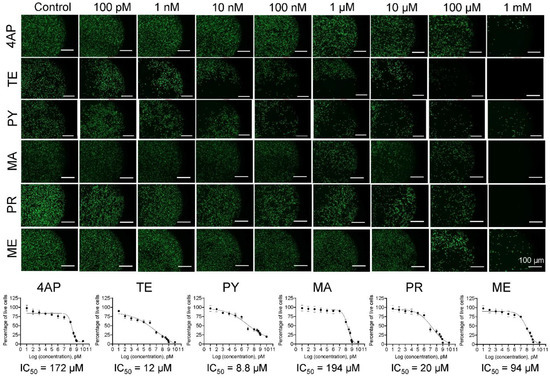
Figure 1
Open AccessReview
Developmental Pattern Formation: Spanish Contributions from a Biophysical Perspective
by
and
Biophysica 2023, 3(2), 335-347; https://doi.org/10.3390/biophysica3020022 - 06 May 2023
Abstract
During the last few decades, developmental pattern formation has evolved from being a descriptive discipline to a quantitative one. That process has been possible due to the implementation of multidisciplinary approaches where biophysicists and mathematicians have played a key role. In this review,
[...] Read more.
During the last few decades, developmental pattern formation has evolved from being a descriptive discipline to a quantitative one. That process has been possible due to the implementation of multidisciplinary approaches where biophysicists and mathematicians have played a key role. In this review, we highlight relevant Spanish contributions and stress their biophysical approaches, as well as provide some historical context. Finally, this work also aimed at bridging the concepts from biology to physics/math (and back) and at shedding light on some directions for future research.
Full article
(This article belongs to the Special Issue State-of-the-Art Biophysics in Spain 2.0)
►▼
Show Figures

Figure 1
Open AccessArticle
Insights into Chemical Interactions and Related Toxicities of Deep Eutectic Solvents with Mammalian Cells Observed Using Synchrotron Macro–ATR–FTIR Microspectroscopy
by
, , , , , , , and
Biophysica 2023, 3(2), 318-334; https://doi.org/10.3390/biophysica3020021 - 04 May 2023
Cited by 1
Abstract
Deep eutectic solvents (DESs) and ionic liquids (ILs) are highly tailorable solvents that have shown a lot of promise for a variety of applications including cryopreservation, drug delivery, and protein stabilisation. However, to date, there is very limited information on the detailed interactions
[...] Read more.
Deep eutectic solvents (DESs) and ionic liquids (ILs) are highly tailorable solvents that have shown a lot of promise for a variety of applications including cryopreservation, drug delivery, and protein stabilisation. However, to date, there is very limited information on the detailed interactions of these solvents with mammalian cells. In this work, we studied six DESs and one IL that show promise as cryoprotective agents, applying synchrotron macro–ATR–FTIR to examine their effects on key biochemical components of HaCat mammalian cells. These data were paired with resazurin metabolic assays and neutron reflectivity experiments to correlate cellular interactions with cellular toxicity. Stark differences were observed even between solvents that shared similar components. In particular, it was found that solvents that are effective cryoprotective agents consistently showed interactions with cellular membranes, while high toxicity correlated with strong interactions of the DES/IL with nucleic acids and proteins. This work sheds new light on the interactions between novel solvents and cells that may underpin future biomedical applications.
Full article
(This article belongs to the Collection Feature Papers in Biophysics)
►▼
Show Figures
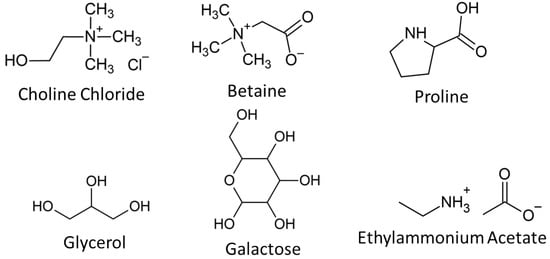
Figure 1
Open AccessArticle
Comparative Investigation of XPS Spectra of Oxidated Carbon Nanotubes and Graphene
by
, , , , and
Biophysica 2023, 3(2), 307-317; https://doi.org/10.3390/biophysica3020020 - 13 Apr 2023
Abstract
X-ray photoelectron emission spectra of thermally reduced graphene oxide samples and carbon nanotubes (CNTs) with various oxidation degrees are presented in this paper. A method for the reconstruction of differential electron inelastic scattering cross sections from the energy loss spectra of photoelectrons is
[...] Read more.
X-ray photoelectron emission spectra of thermally reduced graphene oxide samples and carbon nanotubes (CNTs) with various oxidation degrees are presented in this paper. A method for the reconstruction of differential electron inelastic scattering cross sections from the energy loss spectra of photoelectrons is described and discussed. The analysis of the part of the characteristic photoelectron energy loss spectrum adjacent to the C1 peak indicated a considerable influence of the thermal reduction of graphene oxide on the electron properties of the samples obtained. On the contrary, the oxidation of CNTs by refluxing in a concentrated HNO3 solution does not change the free electron excitation spectrum.
Full article
(This article belongs to the Special Issue Biomedical Optics)
►▼
Show Figures
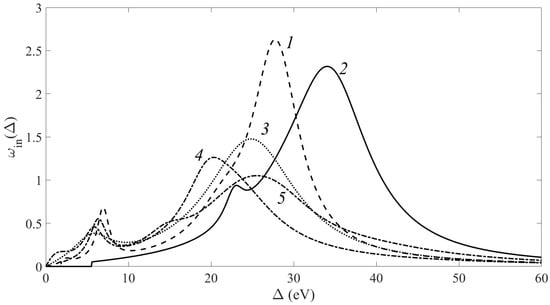
Figure 1
Open AccessArticle
Discovery of the Universal tRNA Binding Mode for the TsaD-like Components of the t6A tRNA Modification Pathway
Biophysica 2023, 3(2), 288-306; https://doi.org/10.3390/biophysica3020019 - 12 Apr 2023
Abstract
Covalent addition of the threonylcarbamoyl group to N(6) of adenosine 37 (t6A modification) within the anticodon loop of several tRNAs is central to the translational fidelity in all known organisms. Structures for each of the enzyme components in the Tsa (t
[...] Read more.
Covalent addition of the threonylcarbamoyl group to N(6) of adenosine 37 (t6A modification) within the anticodon loop of several tRNAs is central to the translational fidelity in all known organisms. Structures for each of the enzyme components in the Tsa (t6A) pathway from all three kingdoms of life have been determined previously. In order to shed light on the poorly defined final step of t6A tRNA modification by TsaD-like components, we performed modeling studies. By docking a tRNA substrate molecule onto reanalyzed complete models of three TsaD-like proteins—TsaD from T. maritima, Qri7 from bacteria, and Kae1 from yeast—we identified a binding site that is common to all of them. An apparently universal binding mode has perfectly oriented tRNA for catalysis by TsaD. Furthermore, it suggests how the conformational changes in TsaD, in response to the binding of the additional regulatory subunits, control enzymatic activity. Re-refinement of the X-ray structure of the TsaBDE complex from T. maritima tentatively suggests that the moiety bound at the active site of the TsaD component is threonylcarbamoyl-AMP (TC-AMP). These findings suggest a detailed model for the mechanism of the catalytic reaction carried out by the TsaD-like components that explains the transfer of unstable TC-AMP from TsaC to TsaD proteins in the t6A modification pathway.
Full article
(This article belongs to the Collection Feature Papers in Biophysics)
►▼
Show Figures
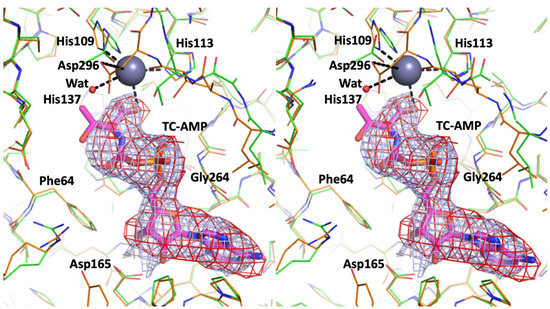
Figure 1
Open AccessArticle
Silica In Silico: A Molecular Dynamics Characterization of the Early Stages of Protein Embedding for Atom Probe Tomography
by
, , , and
Biophysica 2023, 3(2), 276-287; https://doi.org/10.3390/biophysica3020018 - 11 Apr 2023
Abstract
A novel procedure for the application of atom probe tomography (APT) to the structural analysis of biological systems, has been recently proposed, whereby the specimen is embedded by a silica matrix and ablated by a pulsed laser source. Such a technique, requires that
[...] Read more.
A novel procedure for the application of atom probe tomography (APT) to the structural analysis of biological systems, has been recently proposed, whereby the specimen is embedded by a silica matrix and ablated by a pulsed laser source. Such a technique, requires that the silica primer be properly inert and bio-compatible, keeping the native structural features of the system at hand, while condensing into an amorphous, glass-like coating. In this work, we propose a molecular dynamics protocol, aimed at depicting and characterizing the earliest stages of the embedding process of small biomolecules in a solution of water and orthosilicic acid, here, taken as a precursor of the silica matrix. Overall, we observe a negligible influence of orthosilicic acid on the behavior of stable folded systems (such as ubiquitin). Conversely, intrinsically disordered and unstable peptides are affected by the coating, the latter seemingly inhibiting the fluctuations of flexible moieties. While further scrutiny is in order, our assessment offers a first mechanistic insight of the effects of orthosilicic acid, thereby validating its use in the proposed innovative application of APT to the structural resolution of protein molecules.
Full article
(This article belongs to the Collection Feature Papers in Biophysics)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Coarse-Grained MD Simulations of Opioid Interactions with the μ-Opioid Receptor and the Surrounding Lipid Membrane
Biophysica 2023, 3(2), 263-275; https://doi.org/10.3390/biophysica3020017 - 06 Apr 2023
Abstract
In our previous studies, a new opioid (NFEPP) was developed to only selectively bind to the
In our previous studies, a new opioid (NFEPP) was developed to only selectively bind to the
(This article belongs to the Special Issue Molecular Structure and Simulation in Biological System)
►▼
Show Figures
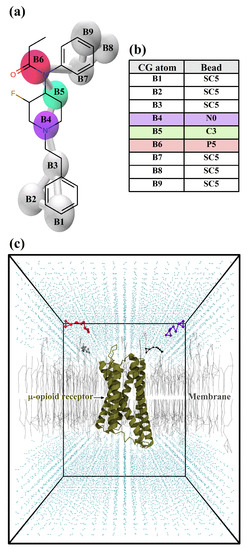
Figure 1
Open AccessArticle
Computational Modeling of the Interaction of Molecular Oxygen with the miniSOG Protein—A Light Induced Source of Singlet Oxygen
Biophysica 2023, 3(2), 252-262; https://doi.org/10.3390/biophysica3020016 - 02 Apr 2023
Abstract
Interaction of molecular oxygen 3O2 with the flavin-dependent protein miniSOG after light illumination results in creation of singlet oxygen 1O2 and superoxide O2●−. Despite the recently resolved crystal structures of miniSOG variants, oxygen-binding sites near the
[...] Read more.
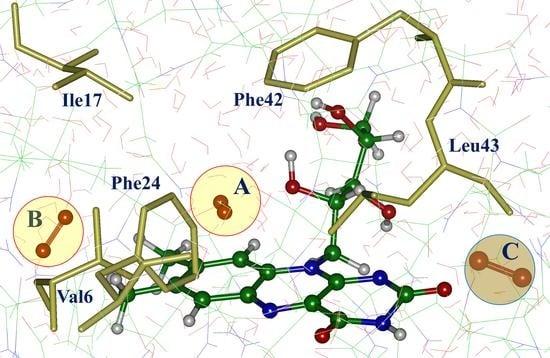
Interaction of molecular oxygen 3O2 with the flavin-dependent protein miniSOG after light illumination results in creation of singlet oxygen 1O2 and superoxide O2●−. Despite the recently resolved crystal structures of miniSOG variants, oxygen-binding sites near the flavin chromophore are poorly characterized. We report the results of computational studies of the protein−oxygen systems using molecular dynamics (MD) simulations with force-field interaction potentials and quantum mechanics/molecular mechanics (QM/MM) potentials for the original miniSOG and the mutated protein. We found several oxygen-binding pockets and pointed out possible tunnels bridging the bulk solvent and the isoalloxazine ring of the chromophore. These findings provide an essential step toward understanding photophysical properties of miniSOG—an important singlet oxygen photosensitizer.
Full article
(This article belongs to the Collection Feature Papers in Biophysics)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Complex Physical Properties of an Adaptive, Self-Organizing Biological System
Biophysica 2023, 3(2), 231-251; https://doi.org/10.3390/biophysica3020015 - 31 Mar 2023
Abstract
Physical modeling of the functioning of the adaptive immune system, which has been thoroughly characterized on genetic and molecular levels, provides a unique opportunity to define an adaptive, self-organizing biological system in its entirety. This paper describes a configuration space model of immune
[...] Read more.
Physical modeling of the functioning of the adaptive immune system, which has been thoroughly characterized on genetic and molecular levels, provides a unique opportunity to define an adaptive, self-organizing biological system in its entirety. This paper describes a configuration space model of immune function, where directed chemical potentials of the system constitute a space of interactions. A mathematical approach is used to define the system that couples the variance of Gaussian distributed interaction energies in its interaction space to the exponentially distributed chemical potentials of its effector molecules to maintain its steady state. The model is validated by identifying the thermodynamic and network variables analogous to the mathematical parameters and by applying the model to the humoral immune system. Overall, this statistical thermodynamics model of adaptive immunity describes how adaptive biological self-organization arises from the maintenance of a scale-free, directed molecular interaction network with fractal topology.
Full article
(This article belongs to the Collection Feature Papers in Biophysics)
►▼
Show Figures

Figure 1
Open AccessArticle
Comparison of Empirical Zn2+ Models in Protein–DNA Complexes
by
and
Biophysica 2023, 3(1), 214-230; https://doi.org/10.3390/biophysica3010014 - 20 Mar 2023
Abstract
Zinc ions are the second most abundant ions found in humans. Their role in proteins can be merely structural but also catalytic, owing to their transition metal character. Modelling their geometric–coordination versatility by empirical force fields is, thus, a challenging task. In this
[...] Read more.
Zinc ions are the second most abundant ions found in humans. Their role in proteins can be merely structural but also catalytic, owing to their transition metal character. Modelling their geometric–coordination versatility by empirical force fields is, thus, a challenging task. In this work, we evaluated three popular models, specifically designed to represent zinc ions with regard to their capability of preserving structural integrity. To this end, we performed molecular dynamics simulations of two zinc-containing protein–DNA complexes, which differed in their zinc coordination, i.e., four cysteines or two cysteines and two histidines. The most flexible non-bonded 12-6-4 Lennard–Jones-type model shows a preference for six-fold coordination of the Zn
(This article belongs to the Collection Feature Papers in Biophysics)
►▼
Show Figures
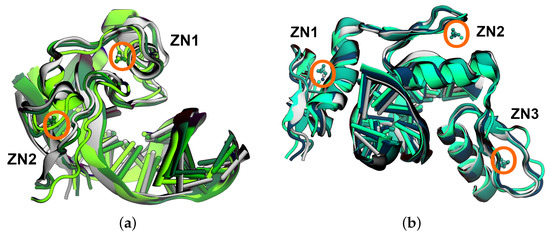
Figure 1
Open AccessArticle
Structural Analysis of Interactions between Epidermal Growth Factor Receptor (EGFR) Mutants and Their Inhibitors
Biophysica 2023, 3(1), 203-213; https://doi.org/10.3390/biophysica3010013 - 14 Mar 2023
Cited by 1
Abstract
People’s lives and health are gravely threatened by non-small-cell lung cancer (NSCLC). Mutations in epidermal growth factor receptor (EGFR), a transmembrane receptor tyrosine kinase, are considered one of the causes of NSCLC. Tyrosine kinase inhibitors (TKIs) are typically used to treat patients with
[...] Read more.
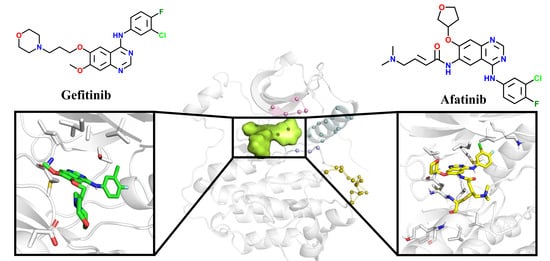
People’s lives and health are gravely threatened by non-small-cell lung cancer (NSCLC). Mutations in epidermal growth factor receptor (EGFR), a transmembrane receptor tyrosine kinase, are considered one of the causes of NSCLC. Tyrosine kinase inhibitors (TKIs) are typically used to treat patients with EGFR mutations. In this study, Gefitinib, a member of the first generation of TKIs, was used to treat an EGFR single-point mutation (single mutant, SM). Patients harboring additional T790M mutations in the kinase domain of the EGFR were resistant to Gefitinib. Then, the L858R/T790M double mutation (double mutant, DM) was treated with the second generation of TKIs, such as Afatinib. Here, we constructed four computational models to uncover the structural basis between EGFR mutants (SM and DM) and corresponding inhibitors (Gefitinib and Afatinib). The binding energy in the G-SM (representing Gefitinib in complex with SM) system was larger than that in the G-DM (Representing Gefitinib in complex with DM) system. Gefitinib’s affinity with L792 and M793 was drastically reduced by the longer side chain of M790 in the G-DM system, which pushed Gefitinib outside of the pocket. Additionally, the A-DM system’s binding energy was higher than the G-DM system’s. Afatinib, unlike Gefitinib, induced the P-loop region to move downwards to decrease the pocket entrance size to accommodate Afatinib properly and stably in the A-DM (Afatinib in complex with DM) system. These results uncover the details of interactions between EGFR and its inhibitors and shed light on the design of new tyrosine kinase inhibitors.
Full article
(This article belongs to the Special Issue Molecular Structure and Simulation in Biological System)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Neuronal Cultures: Exploring Biophysics, Complex Systems, and Medicine in a Dish
Biophysica 2023, 3(1), 181-202; https://doi.org/10.3390/biophysica3010012 - 10 Mar 2023
Abstract
Neuronal cultures are one of the most important experimental models in modern interdisciplinary neuroscience, allowing to investigate in a control environment the emergence of complex behavior from an ensemble of interconnected neurons. Here, I review the research that we have conducted at the
[...] Read more.
Neuronal cultures are one of the most important experimental models in modern interdisciplinary neuroscience, allowing to investigate in a control environment the emergence of complex behavior from an ensemble of interconnected neurons. Here, I review the research that we have conducted at the neurophysics laboratory at the University of Barcelona over the last 15 years, describing first the neuronal cultures that we prepare and the associated tools to acquire and analyze data, to next delve into the different research projects in which we actively participated to progress in the understanding of open questions, extend neuroscience research on new paradigms, and advance the treatment of neurological disorders. I finish the review by discussing the drawbacks and limitations of neuronal cultures, particularly in the context of brain-like models and biomedicine.
Full article
(This article belongs to the Special Issue State-of-the-Art Biophysics in Spain)
►▼
Show Figures
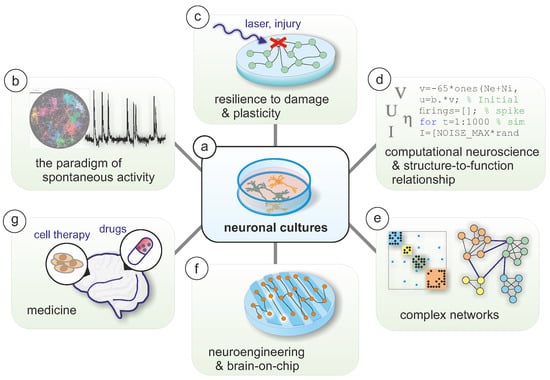
Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Agronomy, Biophysica, IJMS, Life, Plants
Biophysics of Photosynthesis: From Molecules to the Field
Topic Editors: Vasily Ptushenko, Alexei SolovchenkoDeadline: 1 December 2023

Conferences
Special Issues
Special Issue in
Biophysica
State-of-the-Art Biophysics in Spain 2.0
Guest Editor: Jaume CasademuntDeadline: 30 September 2023
Special Issue in
Biophysica
Protein Engineering: The Present and the Future 2.0
Guest Editors: Javier Sancho, Francesca CutruzzolaDeadline: 30 November 2023
Special Issue in
Biophysica
Molecular Structure and Simulation in Biological System 2.0
Guest Editor: Paulino Gómez-PuertasDeadline: 31 December 2023
Topical Collections
Topical Collection in
Biophysica
Feature Papers in Biophysics
Collection Editors: Ricardo L. Mancera, Paul C. Whitford, Chandra Kothapalli